|
7/24/2023 0 Comments Emulsion pcr compared
This is another key value Ion needs to reveal: how much DNA needs to be on a bead to achieve a given read length. In the real world, if a template is larger than this maximum then it will stop amplifying prior to exhaustion of the primers, meaning that the initial signal in sequencing will be lower and will fade to noise sooner.  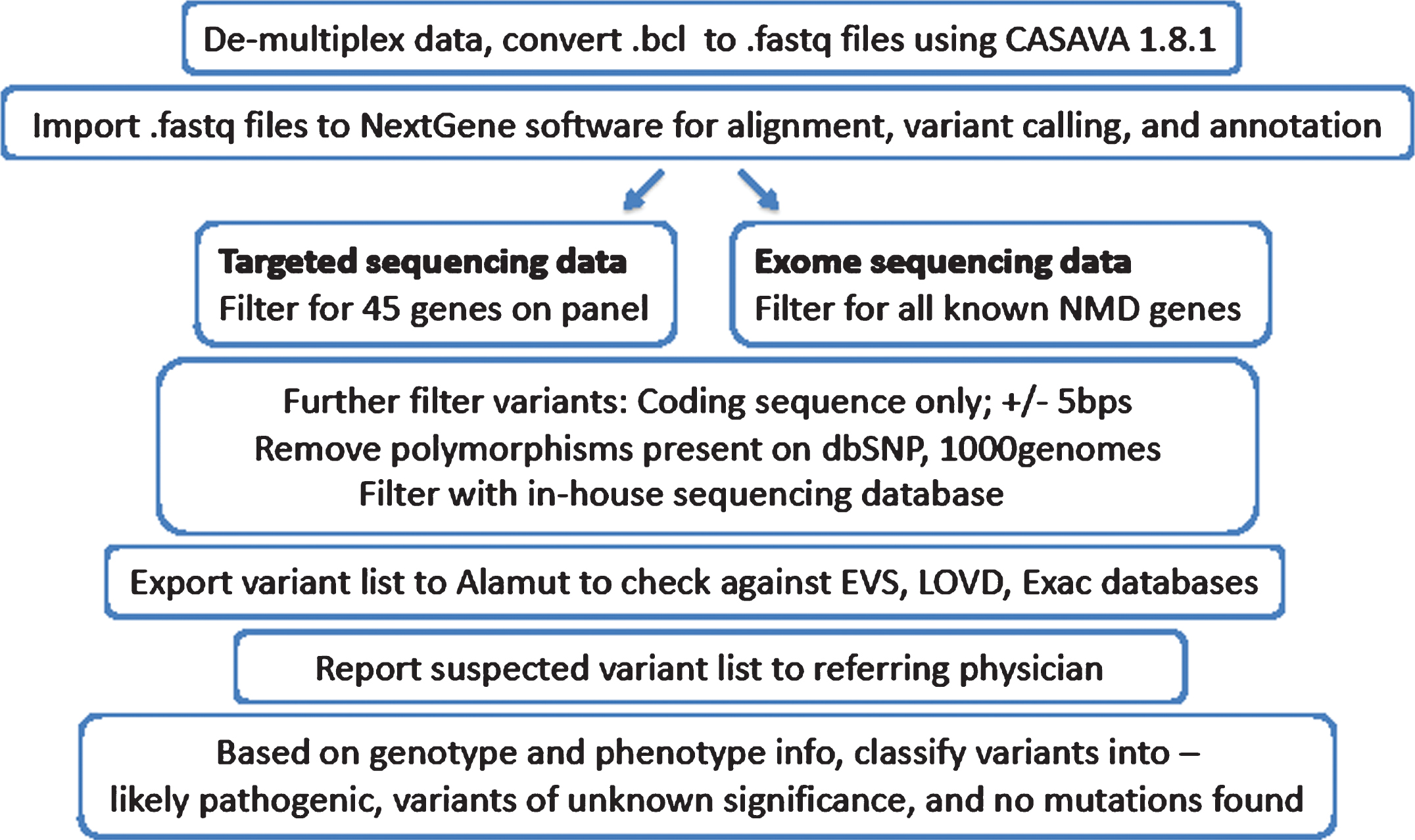
In reality, you would never want the nucleotide concentrations to go anywhere near exhaustion, since the polymerase reaction kinetics will become unfavorable and more errors will be made if the four bases are not depleted uniformly. Clearly, the amount of nucleotides is fixed so the number of nucleotides in the droplet divided by the number of primers is the absolute limit on the length of the amplifiable DNA - to go to completion. Imagine the PCR actually went to completion and consumed all of the primers. This has two important implications.įirst, the size of a droplet and the concentration of the reagents determines the length of the DNA which can be amplified. Please be kind in correcting any math errors I make: I've tried hard to get them right, but this is all stuff I've been able to allow to rust since at least my undergraduate days.Įach droplet has all the reagents which will be available for the PCR run. It's also handy to know that a cubic millimeter is the same as one microliter. Comparing the two required dredging up the formula for the volume of a sphere (4/3 pi r cubed), which I hadn't needed since high school, as well as working with some hairy exponents (by happy chance, TNG is just learning about exponents so I could drive home the point that these are useful, since I'm currently using them!). This is driven by Poisson statistics and is why it is important to know the concentration of a DNA sample going in, in order to set up the most favorable generation of single-template droplets.ĭroplet sizes and beads span a wide range. Well, that's the ideal, though in reality a population of droplets will have some with no DNA, some with one, and some with more than one only the solitary templates will be useful. For second-generation sequencing sample prep, the aqueous droplets also contain a solid bead with at least one primer type bound and also a single initial template DNA molecule. Emulsion PCR (or BEAMing, as one early group called a variant of it), involves generating a water-in-oil emulsion, in which the aqueous phase contains all the components for PCR amplification. Maddeningly, many of the authors in the field tend to publish in journals that I have less than facile access to, but between library visits, PubMed Central and those that are in more accessible journals, I've found a decent start.įirst, the easy stuff. So it is really past time to get more serious about understanding the technology. Plus, there is that temptation to enter the Life Tech grand challenge on sample prep, or attempt to goad some friends into entering. But, I've also seen a 454 experiment go awry in a manner which was blamed on emPCR - a small fraction of primer dimers in our input material became a painful fraction of the sequencing reads. I also have a passing familiarity with RainDance's technology (we participated in the RainDance & Expression Analysis Cancer Grant program). It seems simple in theory, but what goes on in practice? My main reasoning was based on the fact that emPCR is the prep method for both 454 and SOLiD 454 clearly demonstrates the ability to execute long reads (occasionally hitting nearly a kilobase) and SOLiD the ability to generate enormous numbers of beads through emPCR. I've done regular PCR (once in a hotel ballroom, of all places!) but not emulsions. Reading these really drove home to me that I didn't understand emulsion PCR. Some have made rather bold (and negative) predictions, such as Ion Torrent dooming themselves to short read lengths or users being unable to process many samples in parallel without cross-contamination. One theme in some of the comments on my Ion Torrent commentary has been around the limitations of emulsion PCR.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
